Home
Figures
Modules GeneXPress
Global Analysis (Fig. 3) Experimental Tests Candidate Regulators
Technical Reports
Links
People
Supplemental Information
Module Networks: Discovering Regulatory Modules and their Condition Specific Regulators from Gene Expression Data
Much of a cell's activity is organized as a network of interacting functional modules: sets of genes co-regulated to respond to different conditions. We present a probabilistic method for discovering regulatory modules from gene expression data. Our procedure identifies modules of co-regulated genes, their regulators, and the conditions under which regulation occurs, generating testable hypotheses in the form "regulator 'X' regulates process 'Y' under conditions 'W'". We applied the method to a Saccharomyces cerevisiae expression dataset, demonstrating its ability to identify functionally coherent modules and their correct regulators. We present microarray experiments supporting three novel predictions, suggesting regulatory roles for previously uncharacterized proteins. All modules and their associated regulation programs can be viewed here. To examine all modules and regulation programs in more detail, download GeneXPress, and open this file in GeneXPress. 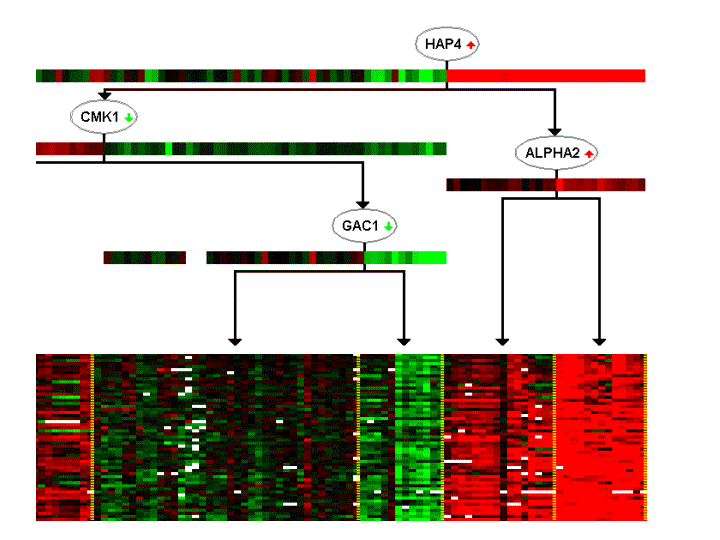 |
||||||||||||||
|
Questions and comments regarding this web site should be addressed to
Eran Segal
Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation. |